Deen Mark G. Toroy*
Armed Forces of the Philippines Medical Center – Victoriano Luna General Hospital Victoriano Luna Avenue, Quezon City, Philippines
*Correspondence: Deen Mark G. Toroy, MD, DPPS, Department of Zoology Armed Forces of the Philippines Medical Center – Victoriano Luna General Hospital Victoriano Luna Avenue, Quezon City, Philippines. E-mail: deentroy@gmail.com
Received: 12 Dec, 2025; Accepted: 22 Dec, 2025; Published: 31 Dec, 2025
Citation: Deen Mark G. Toroy. “Clinical Profile and Selected Viral Genome Sequence Analysis of Sars-Cov-2 Infected Pediatric Patients in A Tertiary Military Hospital in Quezon City from January 2021 – July 2023: A Cross Sectional Study” J Pathol Diagn Microbiol (2025).. 105. DOI: 10.59462/JPDM.1.1.105
Copyright: © 2025 Deen Mark G. Toroy. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Abstract
Background: SARS-CoV-2 displays distinct characteristics in terms of virulence and dis ease severity in pediatric population. Hence, patient’s clinical profile and viral genome may contribute to patient’s symptoms and disease severity
Objectives: To determine the clinical profile and to analyze selected viral genome sequence of pediatric patients infected with SARS-CoV-2 in a Tertiary Military Hospital in Quezon City from January 2021 – July 2023.
Methods: This is a single center cross section al study determining the SARS-CoV infected pediatric patients’ clinical profile consisting of age, sex, residence, comorbidities, presenting symptoms, hospitalization and disease severity. Selected samples were subjected to next gen eration sequencing to analyze the SARS-CoV-2 genome sequence. All data were recorded with utmost confidentiality.
Results & Conclusion: From January 2021 to July 2023, a total of 630 pediatric patients who got tested for SARS-CoV-2 RT-PCR were con firmed to have SARS-CoV-2. Two hundred thir ty-nine (239) were included in this study. Among the study population, 61.51% were male, more than 60% were less than 4 years old and re siding in Quezon City. In terms of clinical pre sentation, 78% had mild symptoms, 20% had radiologic findings of pneumonia, 40% present ed with fever, 12% with diarrhea, and 8% with dyspnea. Ten percent (10%) of patients had concomitant seizure disorder. More than 70% were admitted to the hospital and all of the study subjects recovered from the illness. Only ten samples were subjected to next generation sequencing and all were Omicron variants. Ma jority of the SARS-CoV-2 virus belong to Clade 22B and 23E; and identified pangolin lineages were BA.5.2, BA.2.3.20, XBB.1.5, GJ.1.2 and FL.23.2.
Keywords: SARS-CoV-2, next generation sequencing, viral variants, clinical outcomes
Introduction
The novel coronavirus, SARS-CoV-2, has led to a highly contagious disease that cause a pandemic worldwide. The virus rapidly spreads and evolves as it courses from one continent to the other. It was also observed that several strains of this virus cause different clinical symptoms in different patients. Most notable are the alpha, beta, gamma, delta and omicron variants. Interestingly, infected pediatric population presented different clinical outcomes compared to adults with different strains of virus. [1] During the pandemic, researches on SARS-CoV-2 and their clinical im pact then became an interesting field in epidemiology and infectious disease. There are several published research es on SARS-CoV-2 and their outcomes, likewise there are several published researches on the limited genome se quence analysis of SARS-CoV-2 in the Philippines. How ever, these researches are gathered in the adult population. There is no published research paper yet dedicated to SARS-CoV-2 and their clinical symptoms on pediatric population in the Philippines. Hence, this research was created to investigate the clinical profile of SARS-CoV-2 infected pediatric population and even further look into their viral genome sequences.
Due to the rapid development of technology, specifically in the field of molecular biology, next generation sequenc ing (NGS) was developed in studying viral evolution. The use of NGS technology has become a widely accepted method for outbreak tracking and genomic epidemiology. [1] Detecting new mutations in SARS-CoV-2 allows the researchers to reconstruct unknown routes of infection and provide a molecular basis for SARS-CoV-2 diagnos tic development, vaccine design and drug discovery. [2,3] Hence, viral genomic diversity become an important field in determining viral virulence, transmissibility and disease severity.
There were several researchers worldwide reporting the NGS of SARS-CoV-2 and correlating their clinical symp toms. In the Philippines, coding of SARS-CoV-2 genome was first initiated in August 30, 2020. [4] But due to its high cost and rarity, limited specimen was subjected to NGS. [4] In recent times, there were little researches on SARS CoV-2 genomic sequence in the Philippine settings, [4] more so in the pediatric population. Thus, this research aims to determine the clinical profile of SARS-CoV-2 in fected pediatric patients and analyze selected viral ge nome sequence in a tertiary military hospital in Quezon City. Determining this genome profile will be a vital data that will contribute in the study of SARS-CoV-2 viral evolu tion in pediatric population in the Philippines. Data in this research could be of a great value in future reference, es pecially in the field of epidemiology and vaccine develop ment.
Objectives
General Objectives: To determine the clinical profile and to analyze the selected viral genome sequence of pediat ric patients infected with SARS-CoV-2 in a Tertiary Military Hospital in Quezon City from January 2021 – July 2023.
Specific Objectives:
1. To determine the clinical profile of patients infected with SARS-CoV-2 in a military hospital in Quezon City from January 2021 – July 2023:
a. Age
b. Sex
c. Place of Residence
d. Comorbidities
e. Presenting Symptoms
f. Hospitalization
g. Disease Severity
2. To analyze the SARS-CoV-2 genome sequence of se lected pediatric population using next generation se quencing in a Tertiary Military Hospital in Quezon City from January 2021 – July 2023.
Ethical Consideration
Whole-genome sequencing was performed by collecting the oropharyngeal / nasopharyngeal samples that were collected in the context of routine diagnosis. No addition al samples were collected for this study. Clinical profiles were retrieved from the medical files and anonymized be fore analysis. Ethical approval was obtained from the AFP Medical Center Research Ethics Committee.
Review of Related Literature
During the late December 2019, a novel coronavirus was detected in Wuhan, China. Severe Acute Respiratory Syn drome Coronavirus-2 (SARS-CoV-2) has become a global concern taking millions of lives throughout the globe. [5] The novel virus is capable of rapid transmission through the respiratory epithelium that infects every population that varies from age, race and gender.
This virus is detected through Reverse Transcription Poly merase Chain Reaction (RT PCR) collected from samples of the oral and nasal area of patients. [5] But during the pandemic, the virus evolved, from subvariants that causes minimal symptoms to severe and critical outcomes. Dis ease morbidity and mortality differs from different variants, e.g. the delta variants were more virulent and capable of infecting the pediatric population. [6] Hence, the use of next generation sequencing (NGS), a technology used to investigate the genetic makeup of all living forms capable of sequencing billions of nucleic acid fragments simulta neously has been widely use in studying viral variants. [7] According to the Philippine Pediatric Society (PPS) most children and adolescents infected with SARS-CoV-2 pres ent with mild to moderate symptoms and only a small per centage (0.9%) of patients develop severe and critical dis ease. [8] There are several theories formulated attempting to explain the difference in severity and susceptibility of children compared to adults that includes reduced number of ACE 2 receptors and immature immune systems. [8] However, it was observed that weekly COVID-19 hospi talization rates in children and adolescents have varied throughout the pandemic. [9] Peaking initially in January 2021 (1.5 per 100,000 population), again in September 2021 with predominance of the Delta (B.1.617.2) variant (1.8 per 100,000 population), and then again in Janu ary 2022 with predominance of the Omicron variant (7.1 per 100,000 population). These differences in viral peak opened up a new door of possible different outcomes with different strains of viruses. [9] The viral genomic sequenc ing may pose a great potential in the field of epidemiology and medicine.Infected pediatrics patient’s clinical profile may greatly contribute to their outcome, this include age, sex, comor bidities and their initial presentation. [8] On the contrary, viral strains may not affect the outcome of most patients, but the COVID-19 variants lead to different outcomes manifesting a unique behavior. [9] This may indicate that SARS-CoV-2 viral genomic diversity could be a key in their viral virulence. Like other viruses, SARS-CoV-2 evolves over time. Most mutations in the SARS-CoV-2 genome have no impact on viral function. However, the following notable SARS-CoV-2 variants has different clinical outcomes. [10] The Alpha (B.1.1.7 lineage) was first identi fied in the United Kingdom in late 2020 and subsequent ly became the first globally dominant variant. Alpha was approximately 50 to 75 percent more transmissible than previously circulating strains and it was associated with greater disease severity. [10]
Another is the Beta (B.1.351 lineage), also known as 20H/501Y.V2, was identified and predominated in South Africa in late 2020. [10] The main concern with Beta vari ant was immune evasion: convalescent and post-vaccina tion plasma did not neutralize viral constructs with Beta spike protein as well as those with wild-type spike protein. The Gamma (P.1 lineage), known as 20J/501Y.V3, was first identified in Japan in December 2020. Main concern with this variant is several mutations increasing concern about increased transmissibility and an impact on immu nity. [10]The Delta (B.1.617.2 lineage) lineage was first identified in India in December 2020 and been the most prevalent variant worldwide until emergence of the Omicron variant. Delta variant was more transmissible and was associated with a higher risk of severe disease, hospitalization and death.
COVID 19 in Pediatric Population
During the Omicron dominant periods, weekly hospitaliza tion rates were particularly high in infants younger than six months and in children age six months through four years, who were not eligible for vaccination until June 2022. [11] Among children age 5 to 11 years (who became eligible for vaccination in early November 2021), hospitalization rates were approximately twice as high in unvaccinated than in vaccinated children (19 versus 9 per 100,000) during the period of early Omicron predominance (December 19, 2021 to February 28, 2022). Higher proportions of unvac cinated than fully vaccinated adolescents had COVID-19 as the primary reason for hospitalization (70 versus 41 percent) and required intensive care (30 versus 16 per cent). [11] Several studies support that, COVID-19 disease severity may be related to the strain of SARS-CoV-2. In observa tional studies in children and adolescents, the rates of ad mission to the ICU and mechanical ventilation were lower with the Omicron than the Delta variant. [11] In recent times, the cases of COVID 19 in our country are closely monitored including the variants that is current ly dominant. The use of NGS technology has become a widely accepted method for outbreak tracking and genom ic epidemiology.
Significance of the Study
This is one of the novel studies in the Philippines us ing next generation sequencing (NGS) to determine the SARS-CoV-2 genome sequence infecting pediatric patient in a tertiary military hospital. This paper may strengthen the importance of determining viral genome as a guide in the management and predicting outcomes of COVID-19 infected pediatric patients. This will greatly contribute to the contemporary studies worldwide on SARS-CoV-2 vari ants, the demographics they infect and their clinical symp toms. This can also serve as a literature in other future studies on SARS-CoV-2 genome sequencing and their clinical symptoms. Furthermore, this paper emphasized the importance of NGS and molecular biology in studying viral evolution as a part of diagnostic, therapeutic and vac cine development. In the field of pediatrics, specifically during the COVID-19 pandemic, the viral NGS result was one of the vital data that contribute to the study of the virus. The different viral genome was correlated to the variant, and each variant display distinct characteristics and outcomes. With the aid of NGS, medical personnel become aware of the cir culating virus variant and their behavior. This data helps in planning and formulation of strategies by our medical front-liners, especially during the virulent waves of delta and gamma, wherein healthcare providers become wary and vigilant as cases are expected to be moderate to se vere. On the other hand, during the omicron wave, medical personnel expect a spike in number of infected patients but generally milder cases. With the advent of molecular technology, our understanding of infectious disease be come wider and critical. We can identify the genetic make up of pathogens and correlate it to their virulence. We can develop vaccines congruent to their specific molecular structures. The future of medical research maybe gearing towards understanding the genomic characteristics of the pathogen and its host, and this research is one of those papers pioneering this molecular approach.
Scope and Limitation
This is a single center cross sectional study assessing the clinical symptoms of SARS-CoV- 2 infected pediatric pa tients in a tertiary military hospital. This involves all pediat ric patients seen in V Luna Medical Center from January 2021 – July 2023 that underwent SARS-CoV-2 RT PCR which revealed positive. In addition, selected samples were subjected to next generation sequencing for analy sis. The major limitation of this study is the small number of samples, 10 sequences, hence testing association with other variables were not possible.
Methodology
Study Design
This is a single center cross sectional study determining SARS-CoV-2 genome sequence and patients’ clinical profile in V Luna Medical Center from January 2021 to July 2023. Samples of throat and nasal swabs underwent SARS-CoV-2 RT-PCR and selected samples were subjected to next generation sequencing processed by Armed Forces Research Institute of Medical Sciences (AFRIMS).
Study Population and Setting
Sample population composed of pediatric patients positive for SARS-CoV-2 RT PCR tested in V Luna Medical Center from January 2021 to July 2023. Their demographic and clinical data were retrieved from medical files, including the general data, symptoms, and course of the disease. In addition, selected SARS-CoV-2 RT PCR positive samples with Cycle Threshold of <30 underwent next generation sequencing.
Sample Collection
Nasopharyngeal and oropharyngeal swabs were collect ed using Dacron-tipped swabs by clinical personnel. [12] Samples were collected from pediatric patients under in vestigation for COVID-19, patients seeing clinical care and showing signs of COVID-19-like illness. [12] Swabs were stabilized in various Universal Transport Media (Co pan, CA, USA; Sansure Biotech, Hunan, China; and Sanli, Liuyang, China), temporarily stored at 4°C, transferred to freezers (−80 ± 20°C), and tested within 24 to 72 hours. [12]
SARS-CoV-2 Real-Time RT-PCR Testing Nasopharyngeal and oropharyngeal swabs were collected by clinical staff and rRT-PCR testing was performed. The swabs were heat inactivated at 65°C for 10 minutes and viral RNA was extracted using the NATCH CS automated nucleic acid extraction machine (Sansure Biotech, Hunan, China) or manual RNA extraction kits: QIAmp viral RNA mini kit (Qiagen, Germantown, MD, USA), Sansure RNA one-step nucleic acid release reagent (Sansure, Biotech, Hunan, China), and GenAmplify RNA extraction kit (Ma nila HealthTek, Manila, Philippines) following manufac turers’ instructions. [12] Briefly, specimens were thawed and an aliquot of 200 µL (Sansure) or 140 µL (QIAmp and GenAmplify) of each specimen was used for RNA extraction and eluted in 50 µL following manufacturer’s instructions of each kit. [12] The following SARS-CoV-2 rRT- PCR kits were used following testing conditions and CT cutoff values indicated by each manufacturer: Sansure (Sansure Biotech, Hunan, China), 40 CT; BGI Genomics (Shenzhen, China), 37 CT; modified Berlin assay, 38 CT; and modified U.S. Centers for Disease Control and Pre vention (CDC, USA) assay, 38 CT. The extracted nucleic acid was amplified using the following PCR machines: ABI 7500 (Applied Biosystems, CA, USA), QuantStudio 7 (Ap-plied Biosystems, CA, USA), SLAN96P (Sansure Biotech, Hunan, China), and MA6000 (Sansure Biotech, Hunan, China). [12] Selected specimens also underwent GeneX pert PCR SARS-CoV-2 testing.
SARS-CoV-2 Genome Sequencing and Phy logenetic Analysis
SARS-CoV-2 genome from 10 selected positive speci mens (CT < 30) were sequenced in the in-house molecu lar laboratory, using an iSeq (Illumina, USA), the results of which were reported and described [12] (Table 1).
Phylogenetic Analysis
Phylogenetic analysis was performed using sequences from this study and sequences retrieved from GenBank and GISAID databases on August 24, 2020. [12] The tree was generated using IQ-tree v.1.6.12 using the generalized time-reversible (GTR + F + I) model and 1,000 bootstrap replicates. Multiple sequence alignments were performed using “Multiple Alignment using Fast Fourier Transform” v7.407 with the default setting. [12] The phylogenetic tree was drawn with FigTree v1.4.4 and lineage was deter mined using Pangolin v.2.0.4 (github.com/cov- lineages/ pangolin), including the lineage and the clade of the 23 coding-complete genome sequences from this study. [12] The genome comparisons to the Wuhan-Hu-1 reference genome (GenBank accession number NC_045512.2) were visualized using BRIG v0.95. [12]
Conceptual Framework

Figure 1: Shows the correlation of SARS-CoV-2, patient’s clin ical profile and NGS data
Inclusion Criteria:
1. All pediatric patient 18 years old and below who con sulted in VLMC form January 2021 – July 2023. 2. All pediatric patient who were positive with SARS CoV-2 RT-PCR with and without symptoms. 3. All pediatric patient who were positive for SARS-CoV-2 RT-PCR and underwent next generation sequencing of SARS-CoV-2.
Exclusion Criteria:
1. Non-pediatric age group
2. Positive for SAR-CoV-2 RT-PCR done outside VLMC
Operational Definition of Terms
SARS-CoV-2. The novel coronavirus is the main variable of concern in this paper and includes, determination of its variants and their clinical outcomes in pediatric population.
Reverse Transcription Polymerase Chain Reaction (RT-PCR)
RT-PCR is the diagnostic test for SARS-CoV-2, all sub jects in this paper were positive for SARS- CoV2 RT-PCR before proceeding to NGS.
Next Generation Sequencing (NGS)
Selected subjects in this paper underwent NGS in order to identify the SARS-CoV-2 genomic make-up and viral variants. Qualified samples are those with cycle threshold (CT) value of <30.
Data Collection
Epidemiological, clinical, and disease severity data were extracted from electronic medical records provided by V Luna Medical Center Record Section and Department of Pediatrics Database. All data of patient’s NGS was pro
vided by the AFRIMS. All data records were matched with previously assigned sample IDs and used as anonymized dataset for downstream analyses. All data collected will be treated with utmost confidentiality.
Statistical Interpretation
Measures of central tendency/dispersion and proportion were used to describe quantitative and categorical vari ables.
Results
From January 2021 to July 2023, the Department of Pa thology and Laboratory of V Luna Medical Center tested 103, 486 patients for SARS-CoV-2 RT-PCR, among which 5,945 (5.7%) were pediatric patients. Six hundred thirty (10%) pediatric patients were positive for SARS-CoV-2 and only 10 (1.5%) pediatric patients were subjected to next generation sequencing. (See Figure 2)
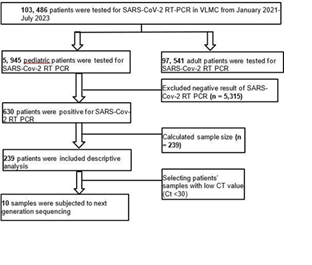
Figure 2: Flow chart of the selection of subjects in the study
A total of 239 patients were included in the study, and ma jority were male 61.51% (n = 147) vs female 38.49% (n = 92) as shown below (Table 1).
|
Gender |
f |
% |
|
Male |
147 |
61.51 |
|
Female |
92 |
38.49 |
Table 1: Shown is the sex distribution of subjects included in the study.
Majority of the subjects were less than 4 years old. One to two years old comprised 27% (n=65), 1mo-11mos 22% (n=53) and 3-4 years old (n=29) as shown in Figure 3 be low.
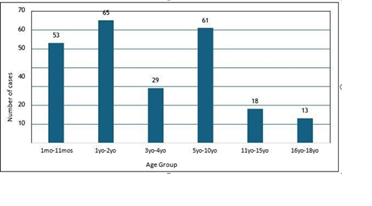
Figure 3: Age distribution of pediatric COVID 19 cases seen in VLMC
Most of the patients were from Quezon City 60.67% (n = 145), followed by Taguig 16.74% (n = 40) and Cavite 15.48% (n = 37) as shown below (Figure 4).
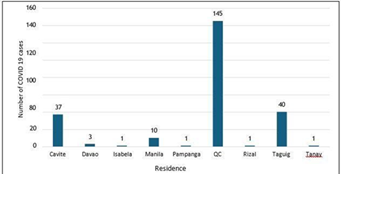
Figure 4: Residence distribution of pediatric COVID 19 cases seen in VLMC.
A quarter of the patients has comorbidities 38% (n = 59) while those with no comorbidities comprised 61% (n = 180) as shown below (Figure 5).
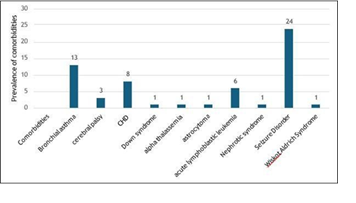
Figure 5: Prevalence of comorbidities among pediatric COVID 19 cases seen in
Most of the patients presented with fever 40% (n = 97) followed by diarrhea 12% (n = 30), cough 9% (n = 22) and dyspnea 8% (n = 21) as shown below (Figure 6). 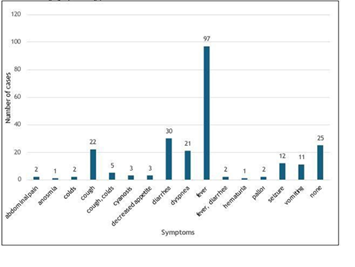 Figure 6: Distribution of symptoms of pediatric COVID 19 cases seen in VLMC.
Figure 6: Distribution of symptoms of pediatric COVID 19 cases seen in VLMC.
Majority of the patients were admitted comprising 76% (n = 183) while 23% (n =56) were advised for home isolation as shown below (Table 2).
|
Hospitalization |
f |
% |
|
Yes |
183 |
76.57 |
|
No |
56 |
23.43 |
Table 2: Shown is the number of subjects who were admitted in the hospital
Most of the patients present with mild symptoms, 78% (n = 187), followed by moderate symptoms 20% (n = 49) and 1% (n = 3) had severe symptoms as shown below (Table 3).
|
Disease Severity |
f |
% |
|
Mild |
187 |
78.24 |
|
Moderate |
49 |
20.50 |
|
Severe |
3 |
1.26 |
Table 3: Shown is the disease severity distribution of subjects included in the study.
More than half of the patients 61% (n = 148) had normal chest X-ray results, while 20% (n = 50) have pneumonia as shown below (Table 4).
|
|---|
Table 4: Shown is the chest x-ray results of subjects included in the study
Out of 239 SARS-CoV-2 positive patients, only ten (10) samples were subjected to NGS Illumina iSeq100TM. NGS was only done for specimen with low cycle threshold value (CT value < 30) that correlates to high viral load (equivalent to ~10,000–100,000 viral genome copies/mL) (Table 7). [7] Genome-wide characterization of SARS-CoV-2 iso lates from 10 samples collected, generated 10 high-quality sequences using iSeq (Illumina, USA) as shown in Table 6. These ten samples revealed Ct value of less than 30:specimen (1) 27.11 – 28.27; specimen (2) 20.78 – 20.68; specimen (3) 15.50; specimen (4) 22.22 – 22.35; specimen (5) 21.68 – 22.10; specimen (6) 16.42 – 16.79; specimen (7) 17.90 – 19.70; specimen (8) 15.79 – 17.90; specimen (9) 27.75 – 34.42 and specimen (10) 19.93 – 20.75.
|
Sample Number |
Obtained consen- sus sequences length (bp)a |
Number of "N" (ambig- uous base) |
Percent reference cover- ageb |
Median depth of coverage (x) |
Nextclade Clade |
Lineage (pango- lin) |
N1 |
Cycle Threshold N2 RP |
|
|
1 |
28829 |
984 |
96.73 |
3557 |
22B (Omicron) |
BA.5.2 |
27.11 |
28.27 |
28.98 |
|
2 |
27814 |
2024 |
94.11 |
1351 |
22B (Omicron) |
BA.5.2 |
20.86 |
20.78 |
31.76 |
|
3 |
28738 |
1102 |
96.98 |
2708 |
22B (Omicron) |
BA.5.2 |
15.50 |
14.99 |
26.29 |
|
4 |
29287 |
555 |
98.47 |
6576 |
21L (Omi- cron) |
BA.2.3.20 |
22.35 |
22.22 |
26.17 |
|
5 |
28247 |
1596 |
95.59 |
1509 |
23A (Omicron) |
XBB.1.5, VOI* |
21.68 |
22.10 |
28.29 |
|
6 |
29354 |
489 |
98.23 |
5806 |
23D (Omicron) |
FL.23.1 |
16.42 |
16.79 |
29.66 |
|
7 |
29360 |
484 |
98.22 |
4458 |
23E (Omicron) |
GJ.1.2 |
19.70 |
17.90 |
27.3 |
|
8 |
23573 |
6263 |
80.90 |
979 |
23A (Omicron) |
XBB.1.5, VOI* |
15.78 |
17.94 |
30.78 |
|
9 |
29352 |
492 |
98.21 |
5818 |
23E (Omicron) |
GJ.1.2 |
27.75 |
34.42 |
32.00 |
|
10 |
23759 |
6084 |
85.81 |
978 |
23E (Omicron) |
GJ.1.2, VUM* |
19.93 |
20.75 |
29.83 |
|
Ave. |
27831.3 |
2,007.3 |
94.3 |
|
|
|
|
|
|
Table 5: Ten SARS COV-2 genome sequence showing the sequences length, ambiguous base, median depth of cover, clades and lineage.
As shown in table 5, Nextclade classified all detected viral strain under Omicron variant. Figure 7 below lifted from Nextstrain.org shows the genomic diversity of SARS CoV-2 identified in this study. [13] Majority of SARS-CoV-2 detected where from the lineages of 22B and 23E.
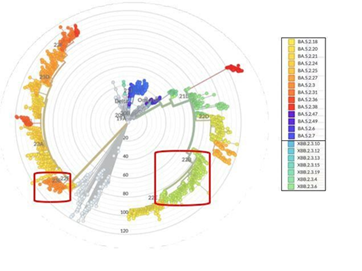
Figure 7: Shown is the complete phylogenetic tree of the novel SARS-CoV-2. [13] Highlighted is the major lineages 22B, 23E and their sub-lineages that were detected in this study. (Image lifted from Nextstarin.org) Furthermore, pangolin lineages identified the following lin eages: BA.5.2, BA.2.3.20, XBB.1.5, GJ.1.2 and FL.23.2 as shown in Table 5.
Specimens 1, 2 and 3 revealed Omicron BA.5.2 lineages, the Illumina run on these 3 clinical specimens obtained consensus sequences length of 28, 829 bp; 27, 814 bp and 28, 738 bp respectively. Mapping against Wuhan SARS-CoV-2 reference (NC_045512) revealed percent reference coverages of 96.73%, 94.11% and 96.98% respectively. These 3 specimens were collected from July to August 2022. Subjects were two males and one female, two were 2-year- old and one is 1-year-old, all were from Quezon City and they all came in due to fever. Two of the subjects were managed as COVID 19 Mild while the other one is managed as COVID 19 Moderate due to the findings of pneumonia on chest x-ray. None of the subjects has comorbidities and all have good clinical outcomes. Specimens 5 and 8 revealed Omicron XBB.1.5 lineage, the Illumina run obtained consensus sequences length of 28, 247 bp and 23, 573 bp respectively. And mapping against Wuhan SARS-CoV-2 reference (NC_045512), the percent reference coverages revealed 95.59% and 80.90% respectively. These specimens were collected last April and May 2023. Subjects were two males, oneis 1-year-old and one is 4-year-old, all were from Quezon City and they came in due to cough, colds and fever. One subject was managed as COVID 19 Mild while the other one is managed as COVID 19 Moderate due to the findings of pneumonia on chest x-ray. One subject has PTB and bronchial asthma, and all subjects have good clinical outcomes. Specimen 7, 9 and 10 revealed Omicron GJ.1.2 lineage, the Illumina run obtained consensus sequences length of 29, 360 bp; 29, 352 bp and 23, 759 bp respectively. Further, mapping against Wuhan SARS-CoV-2 reference (NC_045512) showed percent reference coverages of 98.22%, 98.21% and 85.81% respectively. These 3 specimens were collected from March, May and June 2023. All subjects were female, one is 3-year-old, one is 2-year-old and one is 9-month-old; two is from Quezon City and one is from Tanay, and they all came in due to fever. One of the subjects was managed as COVID 19 Mild while the other two were managed as COVID 19 Moderate due to the findings of pneumonia on chest x-ray. One of the subjects has global developmental delay, hypothyroidism and nephroli thiasis but all have good clinical outcomes.The other 2 specimens revealed unique lineages. Speci men 4 revealed Omicron lineage BA.2.3.20 with percent reference coverage of 98.47%. The specimen was ex tracted from a 3-year- old, male, from Quezon City with bronchial asthma. He was managed as COVID 19 Moder ate due to pneumonia on chest x-ray and was discharged with no complications. On the other hand, the specimen 6 revealed Omicron lineage FL.23.1 with percent refer ence coverage of 98.23%. The specimen was taken from a 4-year-old, male from Quezon City. He was managed as COVID 19 Mild and was discharged with no complications. In terms of demographics and clinical profile, the sex dis tribution of these 10 patients with NGS was 60% (n = 6) males, and 40% (n = 4) females as shown below (Table 6).
|
|---|
Table 6: Shown is the sex distribution of subjects included in the study. Most of the subject in this study are ages 3 years old to 4 years old 40% (n = 4) with mean age distribution of 2.45 years old as shown in Table 7.
|
Age |
f |
|
1mo – 11 mos |
1 |
|
1yo -2yo |
2 |
|
2yo 1mo -3yo |
3 |
|
3yo 1mo – 4yo |
4 |
Table 7: Shown is the age distribution of subjects included in the study.
Majority of the patients are living in Quezon City 90% (n = 9) and 1 subject is outside Metro Manila as shown below (Table 8).
|
Residence |
f |
% |
|
Quezon City, Metro Manila |
9 |
90 |
|
Tanay Rizal |
1 |
10 |
Table 8: Shown are the residence of subjects included in the study.
In terms of comorbidities, 60% (n = 6) have comorbidities in which 2 have bronchial asthma-controlled, pulmonary tuberculosis, umbilical hernia and hydrocele (Table 5). One patient is a 9-month-old with multiple comorbidities: global developmental delay, congenital hypothyroidism and nephrolithiasis as shown below (Table 9).
|
|---|
Table 9: Shown are the number of subjects with comorbidities.
The most prevalent symptoms presented by the subjects were cough 100% (n = 10), followed by fever 70% (n = 7), and colds 20% (n = 2) as shown below (Table 10).
|
Symptoms |
f |
% |
|
Cough |
10 |
100 |
|
Fever |
7 |
70 |
|
Colds |
2 |
20 |
Table 10: Shown is the common symptoms of subjects included in the study.
The severity of COVID 19 was based on the PIDS-P guide lines, [14] wherein 60% (n = 6) were moderate risk due to pneumonia in Chest X-ray (n = 5) and/or concomitant comorbidities (n = 1) as shown below (Table 11 & 12).
|
|---|
Table 11: Shown is the disease severity of subjects included in the study.
|
Chest X-ray |
f |
% |
|
Pneumonia |
5 |
50 |
|
Normal |
3 |
30 |
Table 12: Shown is chest X-ray results of subjects included in the study.
Patients who were tagged COVID-19 moderate were hos pitalized for treatment and close monitoring. None of the patients presented with severe disease and no accounted mortality during the course of the disease as shown below (Table 13).
|
|---|
Table 13: Shown is number of hospitalized subjects included in the study.
Discussion
In this study, we successfully sequenced and character ized ten (10) SARS-CoV-2 samples from COVID-19 pe diatric patients, collected between January 2021 and July 2023, using iSeq (Illumina, USA) in AFRIMS in QuezonCity. All samples collected were Omicron variant. This is congruent to the report of the Philippine Surveil lance and Analysis of COVID-19 in Children Nationwide (SALVACION) database wherein COVID-19 cases in the Filipino pediatric population were Omicron variant and pre sented with mild-to-moderate symptoms (89.5%). [15] In correlation, all patients in this study presented with mild to moderate symptoms and generally had good prognosis and clinical outcomes.
SARS-CoV-2 and Pediatric Age Group Generally, children have milder disease severity than adults. [3] Age becomes significant in terms of vaccine eligibility. [3,6] Several study supports that those with no vaccines are more susceptible to the disease compared to those with vaccines. [3,6,16] In a study by Yigit et al, unvaccinated and incompletely vaccinated children had a higher risk of COVID-19 infection compared to those who were fully vaccinated with an odd ratio of 4.8 and 1.83 re spectively. [16] Since SARS-CoV-2 vaccines were only given to 5 years old and older, those 4 years old and be low are susceptible to contact infection. [3,6] This was as also reflected in this study wherein majority are below 4 years old, 61% (n=147) and were not eligible to receive the vaccine.
SARS-CoV-2, Residence, Sex and Comorbid ities
The current circulating variant becomes the widely detect ed variant in a given period of time and place. In terms of the study setting, the Omicron variant invaded Philippines in the late December 2021, and it was also the most dom inant variant in children. [15] In correlation, all 10 subjects with NGS in this study were infected by Omicron variant. Also, these samples were collected from July 2022 and onwards, hence detecting this variant was generally ex pected. [15] In terms of sex, majority of the subjects in this study were male, 61% (n=147). The difference in sex distribution of the infectious disease is hypothesized to be due to the effect of hormones. [16] In a study by Zaher et al, the estrogen in female enhances T cell activity, stimulates interferon production, increase growth of B cells and antibody pro duction. [16] All of these boost female’s immune system and became a protective mechanism against infection. In contrast, the presence of androgen makes males more susceptible to viral infection. [16] An interesting finding that was recently discovered proposed that expression of TMPRSS2 receptors in peumocytes is androgen-depen dent. [17] TMPRSS2 expression is an important enzyme in SARS-CoV-2 cell entry. [17] Thus, abundant testoster one level in male directly elevates TMPRSS2 expression therefore increasing the probability of viral entry. [17] Lastly, the presence of comorbidities greatly affects the COVID-19 severity. [18,19] This was supported by a me ta-analysis by Tsankov et al and Widjanarko et al, where in those patients with comorbidities have higher risk of developing severe disease with an odd ratio of 1.79 and 4.07 respectively. [18,19] Moreover, studies showed that COVID-19 patient with comorbidities has a higher risk of mortality with relative risk ratio of 2.81. [18] In correlation, in this study 24% (n=12) of patient classified as moderate COVID-19 have preexisting comorbidities, supporting find ing that comorbidities affects patient outcomes and prog nosis.
SARS-CoV-2 Variants and Disease Severity Reports on SARS-CoV-2 revealed that the Gamma and Delta waves lead to severe diseases in children. [20] This was seen in the massive increase in number of admitted children due to COVID- 19 during these 2 waves. On the other hand, Omicron was observed to be more contagious but has lesser disease severity. [22,21] This was also ob served in this study wherein all subjects (n=10) were in fected by Omicron variant and generally have lesser dis ease severity and good clinical outcomes.
Omicron, the most dominant in children The dominance of Omicron variant in pediatric population was theorized to be due to its mode of cellular entry. Ac cording to the PIDS-P COVID-19 guidelines, in contrast to adults, pediatric population are more resistant to former SARS-CoV-2 variants due to children’s innate low amount of TMPRSS2 in type I alveolar epithelial cells, and ACE2 receptors in type II alveolar epithelial cells [14]; and both are vital for viral entry. [14] However, the Omicron variant is unique, that it has higher affinity to ACE2 receptor and solely needs this receptor to invade a host, thus making unvaccinated pediatric patient more vulnerable. [21] Aside from efficient viral entry, the Omicron adapt a more advantageous behavior to spread. A study of Montarrayos et al revealed that Omicron has 6 times higher incidence rate than other variants due to higher effective reproduc tion number (Rt). [22] Interestingly, a study by Chatterjee et. al also shows that Omicron viral load is unique, in that they are concentrated in the upper airway, mostly in the nose, windpipe, and throat, but less in the lower respira tory system. [23] This location makes it easier for the vi rus to be incorporated to respiratory droplets and spread more efficiently. Coupling a faster viral reproduction and concentrated location in the upper respiratory tract, makes omicron more contagious than the previous variants. [23] Lastly, the most important factor in Omicron’s virulence is the viral structure itself. It was proposed by the study of that Omicron’s higher mutation rate compared to oth er variants makes them variant of concern. [24] This was supported by a study of wherein Omicron variant has the highest number of alteration sites, with a staggering 60 substitutions, insertions, and deletions, whereas the sec ond most diverse variant, Delta, has only 22 mutations. [25] This diversity in its protein enables them to take an alternative cell entry pathway (endosomal fusion pathway) and subsequently escaping the host immune system more efficiently. The combined aforementioned characteristics of Omicron could explain why all the subjects in this study are infected by the said variant, as it has more mutations, advantageous behavior to spread and with the most viral load. [25]
Omicron is Less Virulent
But despite its higher infection rate, Omicron infected pa tients had mild to moderate symptoms. As supported by several published studies done in Miami, Ohio and Mexi co, [20,21,22] in comparison to Gamma and Delta variants, Omicron variant caused less severe disease in pediatric population despite its higher incidence rate. Like in this study, all patients presented with cough, colds, fever and loose stool but majority did not require hospital admission. However, there are other contrasting studies on Omicron’s lesser disease severity. A study by Lam et. al., revealed that Omicron caused croup-like symptoms in pediatric pa tient in Hongkong comparing to pre-omicron era leading to more admission cases. [26] It was proposed that the pos sible explanations for greater morbidity is due to its higher transmissibility in younger populations compared with oth er variants [21], along with children’s lack of eligibility for SARS- CoV-2 vaccines, this makes this population more susceptible to omicron. [20] Likewise, a study by Han et. al., also observed that omicron cases had higher peak fe ver and higher rate of croup and febrile seizures compared with the previous waves. [17]
Despite this contrasting studies, strong evidences from a systematic review composed of researches from 52 coun tries worldwide concluded that comparing to Delta variant, Omicron still pales in disease severity. [27] There are no established evidence explaining its lesser disease morbid ity, but a study of Mukherjee et. al hypothesized that this is due to omicron’s low effectivity in counteracting the host cell interferon response. [28] Interferon, a cytokine that plays an important role in the activation of body’s immune defenses. [28] Thus, Omicron’s failure in suppressing in terferon, enables the innate immune system to easily rec ognize and destroy the virus. [28]
Omicron and the Increasing Seizure Inci dence
In comparison to other variants, several studies show that Omicron wave leads to more seizure incident. [29,30] This was also congruent in this study wherein 10% of the pa tient had seizures. Studies of Kim et. al. and Tokuyama et. al., hypothesized that interferon is mainly involved in the increasing incidence of febrile seizure in Omicron wave. [29,30] Because omicron had higher viral replication, this induces higher concentrations of host interferons, a prin cipal component of an innate immune response. 30 This higher amount of interferon then develops to a more in tense innate immune reactions, thus leading to higher grade fever and subsequently higher risk of febrile convul sions. [29,30]
SARS-CoV-2 in Children, the Role of Energy Allocation
There are different researches proposing hypothesis on pediatric resistance to severe COVID-19. Although there are no concrete and established evidences yet, there are theories that became popular in the field of immunology. Since pediatric population manifests different physiology compared to adults, they respond differently in encountering diseases. [31] Human are born defenseless with immature systems, hence, there is a strong evolutionary pressure to mature over time. [31] And one important aspect that is unique in pediatric population, is the rapid growth early in life. [31] This then leads to dilemma of energy allocation trade off, is it whether to invest this energy to produce immune defenses or to prioritize growth? [32] Children’s body will likely favor growth unless the patho gen will lead to life-threatening outcomes. [32] In the study of Brodin, he called this phenomenon, disease tolerance in children, an immune defense strategy activated when the immune response to a pathogen is more damaging than the pathogen itself. [33] The body prevents produc tion and activation of immune response, and tolerates the pathogen rather than resisting it, thereby allocating most of the energy to growth and development. [33] Brodin then concluded that there are higher energy allocation to growth in younger population compared to adult. [33] This is one of the hypothesis explaining why children have lesser COVID-19 disease severity compared to adults, children produce less cytokines and inflammatory agents. [33] Interestingly, male also has higher energy allocation to growth compared to female. [34] This is congruent to our study, wherein 61% (n=147) were less than 4 years old and 61% (n=147) were male, and majority presented with mild symptoms.
Limitation of Next Generation Sequencing Overall, this study was relatively limited by a small number of populations, with just merely ten (10) sequences, this make correlation and drawing conclusions difficult. This limitation is due to the rarity of NGS and its cost effective ness, it is not routinely done, moreso in the developing countries like the Philippines. Despite the small sample, this study supported the findings of other researches that preceded it, a valuable information in understanding the spread and evolution of SARS-CoV-2 in pediatric popula tion in the Philippines. This novel coronavirus was widely studied during the pandemic, but despite the rampant cha os it brought in the community, there were few published literatures of variant outcomes in pediatric population, as well as the aftermath of its rapid evolution.
Conclusion
From January 2021 to July 2023, a total of 630 pediatric patients who got tested for SARS- CoV-2 RT-PCR were confirmed to have SARS-CoV-2. Two hundred thirty-nine (239) were included in this study. Among the study popula tion, 61.51% were male, more than 60% were less than 4 years old and residing in Quezon City. In terms of clinical presentation, 78% had mild symptoms, 20% had radiolog ic findings of pneumonia, 40% presented with fever, 12% with diarrhea, and 8% with dyspnea. Ten percent (10%) of patient had concomitant seizure disorder. More than 70% were admitted to the hospital and all of the study subjects recovered from the illness. Only ten (10) samples were subjected to next generation sequencing and all were Omicron variant. Despite the homogenous variant, the isolated viruses have different lin eages with some tagged as variant of interest and variant under monitoring. Omicron’s dominance may be attributed to its abundant mutation, high effective reproduction num ber and higher affinity to ACE2 receptors compared to oth er variants. Also, majority of the patients presented with mild symptoms such as fever and cough, and generally had good clinical outcomes. This could be associated to Omicron’s inefficient suppression of the host cell interfer on response as opposed to other variants, making it more susceptible to the innate immunity defenses.
Recommendation
Based on the study conclusion, the following recommen dation were deduced:
1. To continue using next generation sequencing as part of work-up especially in complicated, unknown and se vere cases of SARS-CoV-2 and other viral infections.
2. To correlate viral sequence of the SARS-CoV-2 to previous endemic coronavirus in the Philippines and identify mutated sequences that may correlate to its novelty and virulence.
References